| |
13:30
|
0378.
 |
Multi-Band MRSI at 7T using 3D B1 Shimming based Outer Volume
Suppression 
Hoby Patrick Hetherington1, Tiejun Zhao2,
Victor Yushmanov1, and Jullie Pan3
1Radiology, University of Pittsburgh, Pittsburgh,
PA, United States, 2Siemens
Medical Systems, New York, NY, United States, 3Neurology,
University of Pittsburgh, Pittsburgh, PA, United States
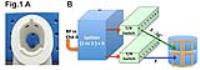
To provide near whole brain coverage for both anatomical
imaging and MRSI we used an 8x2 transceiver array with 8
independent RF channels and eight 1 to 2 splitters. This
configuration provided a homogeneous RF distribution (<12%
SD, 750Hz peak B1) while enabling 3D RF shimming based outer
volume suppression to minimize extra-cerebral lipid signals.
MRSI data was acquired at 7T from control subjects and
patients with mTBI with a multi-band MRSI sequence (four
simultaneous slices) using two RF distributions. Increases
in choline/NAA were seen in both the anterior frontal lobe
and the hippocampi.
|
| |
13:42
|
0379.
 |
Accelerated High-Resolution Multidimensional 1H-MRSI Using
Low-Rank Tensors 
Chao Ma1, Fan Lam1, Qiegen Liu1,
and Zhi-Pei Liang1,2
1Beckman Institute, University of Illinois
Urbana-Champaign, Urbana, IL, United States, 2Electrical
and Computer Engineering, University of Illinois
Urbana-Champaign, Urbana, IL, United States
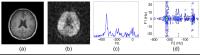
Multidimensional spectroscopy increases spectral dispersion
and enables accurate detection of more metabolites (e.g.,
Glu and GABA in 1H-MRSI of the brain) whose spectra largely
overlap with other metabolites. However, the additional
dimension of spectral information is obtained at the cost of
increased data acquisition time, limiting the practical
utility of in vivo multidimensional MRSI. This work presents
a novel tensor-based approach to accelerated high-resolution
multidimensional 1H-MRSI. The proposed method has been
validated using phantom and in vivo J-resolved 2D 1H-MRSI
experimental studies on a 3T scanner, producing encouraging
results. The method should enhance the practical utility of
multidimensional MRSI.
|
| |
13:54
 |
0380.
 |
Improved spiral chemical shift imaging at 3 Tesla using a
32-channel integrated RF-shim coil array 
Eren Kizildag1, Jason P Stockmann2,
Borjan Gagoski2,3,4, Bastien Guerin2,4,
P. Ellen Grant2,3,4, Lawrence L. Wald2,4,
and Elfar Adalsteinsson1,5,6
1Department of Electrical Engineering and
Computer Science, Massachusetts Institute of Technology,
Cambridge, MA, United States, 2A.
A. Martinos Center for Biomedical Imaging, Massachusetts
General Hospital, Charlestown, MA, United States, 3Boston
Children’s Hospital, Boston, MA, United States, 4Harvard
Medical School, Boston, MA, United States, 5Harvard-MIT
Health Sciences and Technology, Cambridge, MA, United
States, 6Institute
for Medical Engineering and Science, Cambridge, MA, United
States
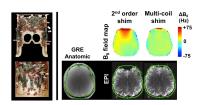
Severe B0 inhomogeneity manifests itself in the
in vivo brain Chemical Shift Imaging (CSI) by broadening the
lineshapes and diminishing the quality of the observed
spectra. We mitigate this problem by employing a 32-channel
integrated RF-shim coil array which uses an optimal
combination of local B0 fields from each coil to
cancel higher order local field inhomogeneities in the CSI
volume. We observed 50% reduction in ΔσB0 over
the slab as compared with 2nd order shimming, corresponding
to pronounced improvements in the linewidths of 13 out of 24
CSI voxels while modestly worsening in only 3 voxels.
|
| |
14:06
 |
0381.
 |
Multi-slice functional FID based spectroscopic imaging on mice
using dynamic shimming at 9.4T 
Aline Seuwen1, Markus Wick2, Franek
Hennel1, Aileen Schroeter1, and Markus
Rudin1,3
1Institute for biomedical engineering, ETH &
University of Zürich, Zürich, Switzerland, 2Bruker
BioSpin MRI GmbH, Ettlingen, Germany, 3Institute
for pharmacology and toxicology, University of Zürich,
Zürich, Switzerland
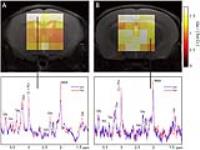
In order to increase the volume coverage of 2D FID based
spectroscopic imaging in mice i.e. the simultaneous
measurement of several brain slices, we implemented a
dynamic shimming approach involving the separate
optimization of first and second order shim terms for
volumes of interest in individual slices. When acquiring two
slices covering cortical and thalamic regions similar
spectra quality has been observed in both slices using
dynamic shimming as compared to measuring each slice
individually. This allows simultaneous acquisition of
metabolite signal changes in several brain regions
associated with stimulus evoked neural activity upon sensory
stimulation.
|
| |
14:18
 |
0382.
 |
Multiband Spectral-Spatial RF Excitation for Hyperpolarized
[2-13C]Dihydroxyacetone 13C-MR Metabolism Studies 
Irene Marco-Rius1, Peng Cao1,
Cornelius von Morze1, Matthew Merritt2,
Karlos X Moreno3, Gene-Yuan Chang4,
Michael A Ohliger1, David Pearce4,
John Kurhanewicz1, Peder EZ Larson1,
and Daniel B Vigneron1
1Department of Radiology and Biomedical Imaging,
University of California San Francisco, San Francisco, CA,
United States, 2Department
of Biochemistry and Molecular Biology, University of
Florida, Gainesville, FL, United States, 3Department
of Chemistry, Engineering, Pre-Pharmacy, and Physics, South
Texas College, Weslaco, TX, United States, 4Department
of Medicine, Division of Nephrology, University of
California San Francisco, San Francisco, CA, United States
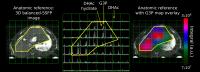
13C-MR spectra of hyperpolarized [2-13C]dihydroxyacetone
(DHAc), a new agent for imaging gluconeogenesis, was
acquired using specialized acquisition methods in the rat
liver and kidney in vivo. Because the resonances originating
from the metabolism of [2-13C]DHAc have a large
frequency distribution, we designed a novel spectral-spatial
(SPSP), multi-band excitation pulse that corrects for
chemical shift misregistration, resulting in accurate
spatial-spectral selectivity. The metabolic products
phosphoenolpyruvate (PEP) and glycerol 3-phosphate (G3P)
were detected, evidencing metabolism of the hyperpolarized
substrate towards the glycolytic pathway and activity of the
enzyme glycerol 3-phosphate dehydrogenase.
|
| |
14:30
 |
0383.
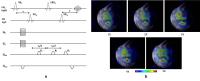 |
Compressed Sensing Accelerated MR Spectroscopic Imaging of
Lactate 
Rohini Vidya Shankar1, Shubhangi Agarwal1,
and Vikram D Kodibagkar1
1Biomedical Engineering, Arizona State
University, Tempe, AZ, United States
Lactate plays a key role in the development and progression
of tumors and its spatial profile can be mapped using
magnetic resonance spectroscopic imaging (MRSI). However,
the long scan time involved in MRSI acquisitions is a
deterrent to its inclusion in routine clinical protocols. A
MRSI sequence containing lactate editing components combined
with prospective compressed sensing acquisitions was
developed for fast mapping of lactate metabolism,
particularly in response to treatment. Results from in vivo
experiments demonstrate a reduction in acquisition time by
up to 80%, with the accelerated MRSI datasets maintaining
high fidelity with the fully sampled reference dataset.
|
| |
14:42
 |
0384.
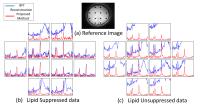 |
Low-rank based compartmentalized reconstruction algorithm for
high resolution MRSI without lipid suppression methods 
Ipshita Bhattacharya1 and
Mathews Jacob1
1Department of Electrical and Computer
Engineering, The University of Iowa, Iowa City, IA, United
States
A novel compartmental low rank algorithm and data
acquisition method for high resolution MR spectroscopic
imaging without the use of any lipid suppression methods is
introduced. The field inhomogeneity compensated data is
modeled as the sum of a lipid dataset and a metabolite
dataset using the spatial compartmental information obtained
from the water reference data. These datasets are modelled
to be low-rank subspaces and are assumed to be mutually
orthogonal. The high resolution spiral acquisition method
achieves in plane resolution of upto 1.8x1.8 mm2 in
7.2 mins. Recovery from these measurements is posed as a low
rank recovery problem. Experiments on in-vivo data
demonstrates comparable results for both lipid suppressed
and lipid unsuppressed data.
|
| |
14:54
 |
0385.
 |
Ultrahigh-Resolution Metabolic Imaging at 9.4 Tesla 
Fan Lam1, Hanbing Lu2, Yihong Yang2,
Bryan Clifford1,3, Chao Ma1, Gene E
Robinson4, and Zhi-Pei Liang1,3
1Beckman Institute, University of Illinois at
Urbana-Champaign, Urbana, IL, United States, 2Neuroimaging
Research Branch, National Institute on Drug Abuse,
Baltimore, MD, United States, 3Department
of Electrical and Computer Engineering, University of
Illinois at Urbana-Champaign, Urbana, IL, United States, 4Carl
R. Woese Institute for Genomic Biology, University of
Illinois at Urbana-Champaign, Urbana, IL, United States
We present a multislice short-TE 1H-MRSI method to achieve
fast, ultrahigh-resolution metabolic imaging of rats on a
9.4 Tesla animal scanner. The proposed method uses a
subspace-based hybrid data acquisition strategy and a
low-rank-model-based image reconstruction scheme. In vivo
experiments have been performed to demonstrate the
feasibility of the proposed method. We are able to produce
high-SNR, spatially resolved metabolic profiles from the rat
brain with 1x1x2mm3 nominal
resolution in 16 minutes.
|
| |
15:06
|
0386.
 |
Overdiscrete Reconstruction in Echo-Planar Spectroscopic
Imaging with Auto Calibrated B0 Field Map Estimation 
Eduardo Coello1,2, Martin Janich2,
Timo Schirmer2, Ralf Noeske3, Tamas
Borbath2, Axel Haase1, and Rolf
Schulte2
1Technische Universität München, Munich, Germany, 2GE
Global Research, Garching, Germany, 3GE
Healthcare, Potsdam, Germany
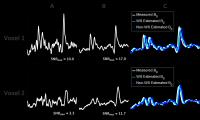
An overdiscrete reconstruction for in-vivo 3D Echo-Planar
Spectroscopic Imaging (EPSI) data is used for SNR
improvement and voxel bleeding reduction. We propose the
estimation of a B0 field
map, which is needed for the reconstruction, using the
residual water signal in the dataset. A mean SNR enhancement
of a factor of 2.8 was achieved for NAA and comparable
reconstruction results were obtained with both the measured
and the estimated B0 field
maps.
|
| |
15:18
|
0387.
 |
Metabolic mapping of the brain using ultra-high resolution MRSI
at 7 T 
Gilbert Hangel1, Bernhard Strasser2,
Michal Považan2, Lukas Hingerl1, Marek
Chmelík2, Stephan Gruber2, Siegfried
Trattnig2,3, and Wolfgang Bogner2
1MR Centre of Excellence, Medical University of
Vienna, Vienna, Austria, 2MRCE,
Department of Biomedical Imaging and Image-guided Therapy,
Medical University of Vienna, Vienna, Austria, 3Christian
Doppler Laboratory for Clinical Molecular MR Imaging,
Vienna, Austria
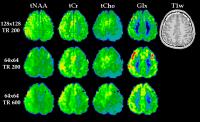
Increasing the resolution of MRSI is desirable to delineate
small structures and pathologic deviations such as Multiple
Sclerosis lesions and increase local B0-homogeneity per
voxel. We show that using an FID-MRSI sequence with short TR
and L2-regularisation for lipid contamination removal, the
major brain metabolites can be mapped with a 128x128 matrix
over a whole brain slice with unprecedented detail, with a
nominal voxel volume of 1.7×1.7×8 mm³. The additional
application of parallel imaging allows reducing measurement
times enough for potential clinical applications.
|
|












