|
|
|
Plasma # |
|
0001.
 |
1 |
DWI^2: exploring the MRI-phase for imaging diffusion 
Ralph Sinkus1, Simon Auguste Lambert1,
Lucas Hadjilucas1, Shaihan Malik2,
Anirban Biswas1, Francesco Padormo2,
Jack Lee1, and Joseph V Hajnal2
1Imaging Sciences & Biomedical Engineering
Division Kings College, King's College London, London,
United Kingdom, 2Centre
for the Developing Brain & Department Biomedical
Engineering, King's College London, London, United Kingdom
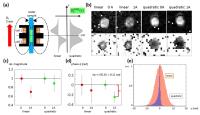
Classical DWI methods extract information about
microstructural tissue complexity from the signal decrease
of the MR-magnitude as a function of b-value. Utilization of
linear gradients for motion encoding prevents theoretically
the use of the MR-phase. Rather, the diffusion information
is encoded in the MR-magnitude via global spin dephasing due
to Brownian motion with zero net phase shift. This dogma is
overturned when considering quadratic gradient fields in
space. We demonstrate in theory, experiment, and simulation
that the diffusion process leads to a net phase shift with
minimal loss in signal magnitude when imaging at the minimum
of the quadratic gradient.
|
|
0002.
 |
2 |
High resolution diffusion tensor reconstruction from
simultaneous multi-slice acquisitions in a clinically feasible
scan time 
Gwendolyn Van Steenkiste1, Ben Jeurissen1,
Steven Baete2,3, Arnold J den Dekker1,4,
Dirk H.J. Poot5,6, Fernando Boada2,3,
and Jan Sijbers1
1iMinds-Vision Lab, University of Antwerp,
Antwerp, Belgium, 2Center
for Advanced Imaging Innovation and Research (CAI2R), NYU
School of Medicine, New York, NY, United States, 3Center
for Biomedical Imaging, Department of Radiology, NYU School
of Medicine, New York, NY, United States, 4Delft
Center for Systems and Control, Delft University of
Technology, Delft, Netherlands, 5Imaging
Science and Technology, Delft University of Technology,
Delft, Netherlands, 6Biomedical
Imaging Group Rotterdam, Erasmus Medical Center Rotterdam,
Rotterdam, Rotterdam, Netherlands
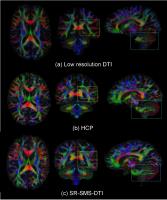
Achieving a high spatial resolution with DTI is challenging
due to the inherent trade-off between resolution,
acquisition time and signal-to-noise ratio (SNR). We propose
a strategy to improve this trade-off by combining
super-resolution DTI (SR-DTI) and simultaneous multi-slice
(SMS) acquisition. With SMS-SR-DTI, high resolution DTI
parameters can be recovered from thick slice images which
have a high SNR. By acquiring the images with SMS, the
overall acquisition time remains clinically feasible. As
such, high resolution in vivo DTI becomes feasible in a
clinical setting. This opens up exciting possibilities for
diffusion MRI research.
|
 |
0003.
 |
3 |
Quantitative evaluation of eddy-current and motion correction
techniques for diffusion-weighted MRI 
Mark S Graham1, Ivana Drobnjak1, and
Hui Zhang1
1Centre for Medical Image Computing & Department
of Computer Science, UCL, London, United Kingdom
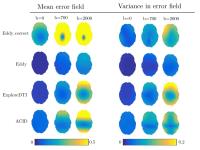
It is necessary to perform correction of eddy-current and
motion artefacts before analysing DW-MR data, but none of
the commonly used correction techniques have been evaluated
quantitatively using direct measures of correspondence. Here
we apply a recently proposed simulation framework to
evaluate four correction techniques. We found the three
techniques that register to a b=0 image (Eddy_correct, ACID,
ExploreDTI) perform worse than a technique that registers to
predicted DWIs (eddy). Furthermore, we found that one of
the most commonly used methods for registration to b=0,
eddy_correct, performs significantly worse than the other
methods considered.
|
|
0004.
 |
4 |
A Mathematical Model and an Efficient Simulation Framework for
Diffusion Cardiac Imaging: Application to Quantification of
Cardiac Deformation on the Diffusion Signal 
Imen Mekkaoui1, Kévin Moulin2,3,
Jérôme Pousin1, and Magalie Viallon2,4
1ICJ, INSA-Lyon, Villeurbanne, France, 2Creatis,
INSA-Lyon, Lyon, France, 3Siemens
Healthcare, Saint-Denis, France, 4Department
of Radiology, Universite´ J. Monnet, Saint Etienne, France
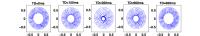
The diffusion process in the myocardium is difficult to
investigate because of the unqualified sensitivity of
diffusion measurements to cardiac motion. We introduced a
mathematical formalism to quantify the effect of tissue
motion on the diffusion NMR signal. The presented model is
based on the Bloch-Torrey equations and takes into account
the cardiac deformation according to the laws of continuum
mechanics. Approximating this model by using a finite
element method, numerical simulations can predict the
sensitivity of the signal to cardiac motion under the
influence of different preparation schemes. Our model
identified the existence of two time points of the cardiac
cycle, called "sweet spots", on which the diffusion is
unaffected by the cardiac deformation. This study also
demonstrates that the sweet spots depend on the type of
diffusion encoding schem.
|
 |
0005.
 |
5 |
Diffusion Kurtosis at varying diffusion times in the normal and
injured mouse brains 
Dan Wu1, Frances J Northington2, and
Jiangyang Zhang1,3
1Radiology, Johns Hopkins University School of
Medicine, BALTIMORE, MD, United States, 2Pediatrics,
Johns Hopkins University School of Medicine, BALTIMORE, MD,
United States, 3Radiology,
New York University School of Medicine, New Yourk, NY,
United States
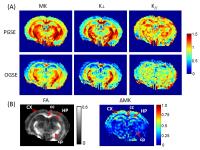
To investigate the diffusion time dependence of diffusion
kurtosis, we measured kurtosis at varying diffusion times
using pulsed and oscillating gradients. The results showed
reduced kurtosis as diffusion time decreased from 25 ms to
2.5 ms in the normal adult mouse brains, and the differences
were higher in the gray matter than the white matter
regions. Results from neonatal mice with severe
hypoxic-ischemic injury showed that both kurtosis
measurements at short and long diffusion times elevated in
the edema region, and the changes were heterogeneous in the
hippocampus, which may be correlated with long-term
outcome.
|
|
0006.
 |
6 |
Can the Stretched Exponential Model of Gas Diffusion
Provide Clinically -Relevant Parenchyma Measurements of Lung
Disease? 
Alexei Ouriadov1, Eric Lessard1, David
G McCormack2, and Grace Parraga1
1Robarts Research Institute, The University of
Western Ontario, London, ON, Canada, 2Department
of Medicine, The University of Western Ontario, London, ON,
Canada
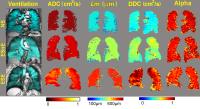
We hypothesized that using inhaled noble gas MRI
diffusion-weighted imaging, the diffusion scale estimated
using the stretched exponential model would be strongly
related to MRI estimates of the mean linear intercept of the
lung parenchyma. In this proof-of-concept evaluation, we
evaluated 34 never- and ex-smokers and compared parenchyma
morphological estimates acquired using two different MRI
approaches ad as well with CT and pulmonary function test
measurements of acinar duct structure and function. This is
important because in obstructive lung disease, the
non-invasive measurement of parenchyma tissue destruction or
maldevelopment may serve as a therapeutic target.
|
|
0007.
 |
7 |
Overestimation of CSF fraction in NODDI: possible correction
techniques and the effect on neurite density and orientation
dispersion measures 
Samira Bouyagoub1, Nicholas G. Dowell1,
Samuel A. Hurley2, Tobias C. Wood3,
and Mara Cercignani1
1Clinical Imaging Sciences Centre, Brighton and
Sussex Medical School, Brighton, United Kingdom, 2FMRIB
Centre, University of Oxford, Oxford, United Kingdom, 3Neuroimaging,
IoPPN, King’s College London, London, United Kingdom
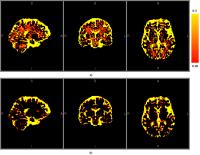
NODDI is a diffusion MRI technique based on combining a 3
compartment tissue model with a (HARDI) protocol. NODDI
overestimates CSF volume fractions (fiso), particularly in
white matter regions. This is possibly due to the single T2
assumption for all compartments. High fiso could lead to
inaccurate measure of neurite density (ficvf) and
orientation dispersion (odi). We propose a method to correct
these errors by scaling fiso with voxel-based T2 maps from
DESPOT. We acquired NODDI data for 5 healthy subjects, and
we run original NODDI analysis and another NODDI analysis
using rescaled fiso. Results showed rescaling fiso generated
low fiso measures consistent with those reported in
literature. It also generated more physiologically
acceptable measures of ficvf, whereas odi was not sensitive
to the change.
|
|
0008.
 |
8 |
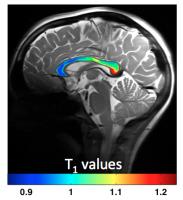
Quantitative Assessment of Microstructure Properties of Human
Corpus Callosum and Distinct Connectivity to Projected Cortices
using Parametric T1 Imaging and Diffusion Tractography - Permission Withheld
Byeong-Yeul Lee1, Xiao-Hong Zhu1, and
Wei Chen1
1Center for Magnetic Resonance Research,
Radiology, University of Minnesota, Minneapolis, MN, United
States
Imaging of callosal microstructures is of importance to
understand its functional and anatomical connectivity to the
projected cortical areas across two hemispheres. In this
work, we tested our hypothesis that the parametric T1 measure
could be sensitive to the corpus callosum (CC)
microstructure and the fiber size within CC, and it may
reflect the underlying functionality. In comparison with
histology reports, our T1 maps
indicate high inhomogeneity in CC and a positive trend
between the T1 value
and CC fiber size. In addition, diffusion tractograpy
analysis shows that regional differentiation of CC T1 value
or fiber size is indicative of unique connection to the
cortical areas with distinct brain function. We found that
the large callosal fibers likely connect to sensory and
visual cortices; in contrast, small callosal fibers connect
higher functional brain regions. The overall results show
the new utility of parametric T1 imaging
for quantitatively assessment of the fiber microstructure of
human corpus callosum and its connections to functionally
relevant cortices. This imaging approach could provide a
robust and useful tool for detection of fiber abnormality in
the human white matter and dysfunction.
|
 |
0009.
 |
9 |
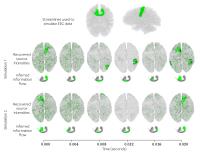
Fibre directionality and information flow through the white
matter: Preliminary results on the fusion of diffusion MRI and
EEG 
Samuel Deslauriers-Gauthier1, Jean-Marc Lina2,
Russell Butler3, Kevin Whittingstall3,
Pierre-Michel Bernier4, and Maxime Descoteaux1
1Computer Science department, Université de
Sherbrooke, Sherbrooke, QC, Canada, 2École
de Technologie Supérieure, Montréal, QC, Canada, 3Department
of Diagnostic Radiology, Université de Sherbrooke,
Sherbrooke, QC, Canada, 4Department
of Kinanthropology, Université de Sherbrooke, Sherbrooke,
QC, Canada
Diffusion MRI can recover white matter fibre bundles but it
is blind to their directionality. We propose to identify the
directionality of white matter fibre bundles by combining
diffusion MRI and EEG data. Based on a realistic model of
the brain and simulated EEG data, our preliminary results
show that our proposed method is able to differentiate
between afferent and efferent white matter connections.
|
 |
0010.
 |
10 |
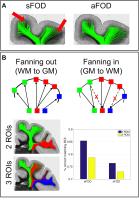
Improved tractography by modelling sub-voxel fibre patterns
using asymmetric fibre orientation distributions 
Matteo Bastiani1, Michiel Cottaar1,
Krikor Dikranian2, Aurobrata Ghosh3,
Hui Zhang3, Daniel C. Alexander3,
Timothy Behrens1, Saad Jbabdi1, and
Stamatios N. Sotiropoulos1
1FMRIB Centre, University of Oxford, Oxford,
United Kingdom, 2Department
of Anatomy & Neurobiology, Washington University, St. Louis,
MO, United States, 3Department
of Computer Science & Centre for Medical Image Computing,
University College London, London, United Kingdom
Fiber bundles can cross or kiss, bend or fan within a single
diffusion MRI (dMRI) voxel. Given the limited dMRI
resolution and the inherent central symmetry in the
measurements, these sub-voxel patterns cannot be
distinguished by only using the voxel-wise signal. These
asymmetric fibre patterns can be distinguished once
information from neighbouring voxels is pooled together. We
propose a direct estimation of asymmetric fiber orientation
distributions (aFODs) based on neighbourhood-wise constrained
spherical deconvolution that is capable of inferring
sub-voxel patterns. We also propose a tractography algorithm
based on the estimated aFODs and we assess performance using
real histological fibre patterns.
|
|
0011.
 |
11 |
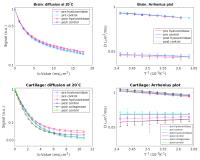
Investigation of the influence of the extracellular matrix on
water diffusion in brain and cartilage 
Jakob Georgi1, Riccardo Metere1,
Markus Morawski2, Carsten Jäger2, and
Harald E. Möller1
1Max-Planck-Institute for Human Cognitive and
Brain Sciences, Leipzig, Germany, 2Paul-Flechsig-Institute
for Brain Research, Leipzig, Germany
Water diffusivity in biological tissues can be related to
the underlying microstructure that modulates the restricted
or hindered diffusion, and can be studied with NMR
experiments. The extracellular matrix, whose composition
depends on the tissue type, may have an influence on
diffusion. In this work we study the influence of the
extracellular matrix on diffusion, by measuring brain and
cartilage samples before and after the enzymatic removal of
the extracellular matrix components. The activation energy
for the self-diffusion of water seems to be not
significantly affected by the treatment for brain tissues.
|
|
0012.
 |
12 |
Measurement of the Effect of Tissue Fixation on Tumour
Microstructure using VERDICT Diffusion-MRI 
Ben Jordan1, Tom Roberts1, Angela
D'Esposito1, John Connell1, Andrada
Ianus2, Eleftheria Panagiotaki2,
Daniel Alexander2, Mark Lythgoe1, and
Simon Walker-Samuel1
1Centre for Advanced Biomedical Imaging,
University College London, London, United Kingdom, 2Centre
for Medical Image Computing, University College London,
London, United Kingdom
It has previously been shown that compartmental models of
tissue diffusion such as VERDICT can enable access to useful
measures of in-vivo tumour microstructure such as cell
radius. However, comparing the in-vivo values with those
measured from histology showed that a discrepancy exists
between the two; histological values were consistently
smaller. In this study, we assess the ability of VERDICT MRI
to detect this change in cell radius by acquiring data (9.4T
MRI) both in-vivo and post-fixation. A significant decrease
in cell radius was detected post-fixation, which was
supported by a decrease in the intra-cellular volume
fraction.
|
 |
0013.
 |
13 |
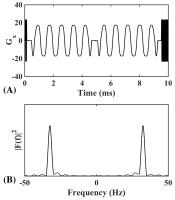
Validation of Surface-to-Volume Ratio derived from Oscillating
Gradient Spin Echo on a clinical scanner using anisotropic fiber
phantoms 
Gregory Lemberskiy1, Steven H. Baete1,
Martijn A. Cloos1, Dmitry S. Novikov1,
and Els Fieremans1
1Radiology, NYU School of Medicine, New York, NY,
United States
This work represents the first measurement of S/V on a
clinical scanner using OGSE on a well-characterized
anisotropic fiber phantom. The S/V measurement is validated
externally via non-diffusion metrics (Spin Echo and MR
Fingerprinting). Lastly, a comparison of $$$D(\omega)$$$ and
$$$D(t)$$$ shows that the effective diffusion time is
$$$t_{\rm eff}^{\rm Mitra} = 9/64f = 9/16\cdot t_{\rm
eff}$$$ rather than the commonly used $$$t_{\rm eff} =
1/4f$$$.
|
 |
0014.
 |
14 |
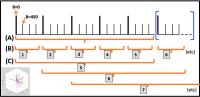
Demonstration of a Sliding-Window Diffusion Tensor Technique for
Temporal Study of Post-Exercise Skeletal Muscle Dynamics 
Conrad P Rockel1,2 and
Michael D Noseworthy1,2,3
1School of Biomedical Engineering, McMaster
University, Hamilton, ON, Canada, 2Imaging
Research Centre, St Josephs Healthcare, Hamilton, ON,
Canada, 3Electrical
and Computer Engineering, McMaster University, Hamilton, ON,
Canada
A novel sliding-window DTI analysis strategy, aimed at
achieving both temporal resolution and valid spatial
representation, was tested on one human subject pre- and
post-exercise (plantar flexion) across 4 sets of different
intensity. Temporal diffusion measures comprised of 3- and
15-directions (ADC and MD/FA, respectively) were assessed,
as well as signal intensity of accompanying T2-weighted
images (S0). Peroneus longus demonstrated increase in MD,
ADC and S0, the peak and duration of which reflected
exercise intensity. FA appeared noisy, although
demonstrated large decreases following higher intensity
exercise. While further work is needed, this method shows
promise in measuring skeletal muscle dynamics.
|
|
0015.
 |
15 |
Denoising Diffusion-Weighted Images Using x-q Space Non-Local
Means 
Geng Chen1,2, Yafeng Wu1, Dinggang
Shen2, and Pew-Thian Yap2
1Data Processing Center, Northwestern
Polytechnical University, Xi'an, China, People's Republic
of, 2Department
of Radiology and BRIC, University of North Carolina at
Chapel Hill, Chapel Hill, NC, United States
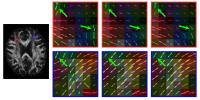
In this abstract, we show that improved denoising
performance can be attained by extending the non-local means
(NLM) algorithm beyond the x-space (i.e., the
spatial space) to include the q-space
(i.e., the wave-vector space). The advantage afforded by
this extension is twofold: (1) Non-local information can now
be harnessed not only across space, but
also across measurements in q-space;
(2) In white matter regions with high curvature, q-space
neighborhood matching corrects for such non-linearity so
that information from structures oriented in different
directions can be used more effectively for denoising
without introducing artifacts.
|
|

















