ISMRM 24th Annual Meeting & Exhibition • 07-13 May 2016 • Singapore |
|
Weekend Educational Course: Advanced MR Spectroscopy in Operation
Skill Level: Intermediate
Organizers: Anke Henning, Ph.D. & Carolyn E. Mountford, D.Phil.(Oxon)
Sunday 08 May 2016 |
Overview
Advanced MRS methods such as whole organ 1H and 31P MRSI, spectral
editing and 2D MRS will be presented that recently enabled novel
clinical and research applications in the fields of brain (psychiatric
disorders, traumatic brain injury, PTSD, pain); breast (prognosis in
high risk woman) and prostate (prognostic cancer imaging) diseases.Target Audience
Psychiatrists, Neurologists, Neuroradiologists, Body Radiologists, MR
Physicist who support this groups, Ph.D. students and Postdoctoral
fellows in related research areas.
Educational Objectives
Upon completion of this course, participants should be able to:
- Understand the basic
principles of advanced MRS methods;
- Understand the added value of
advanced MRS methods; and
- Understand which clinical fields
benefit from advanced MRS methods.
|
|
PROGRAM |
| Moderators: Anke Hennig, Carolyn Mountford |
| |
|
|
MRSI |
|
13:30
|
|
Basic Principles & Sequences for Whole Organ MRSI - Brain &
Body 
Dennis W. J. Klomp1
1UMC Utrecht
In this course, the basics of maximizing SNR while
minimizing sensitivity to system imperfections in MRSI
of the human body are discussed using example
applications in brain, prostate, breast, and body tuned
for the nucleus of 1H, 31P
and 19F.
|
13:50
|
|
Whole Brain (Organ) MRSI Analysis - Permission Withheld
Peter B Barker1
1Radiology, Johns Hopkins Univ School of
Medicine, Baltimore, MD, United States
The efficient and accurate analysis of whole brain MR
spectroscopic imaging (MRSI) data is critical for the
acceptance of this technique into both research and
clinical usage. This presentation will review methods
for the processing of MRSI data collected with extended
brain coverage and high spatial resolution, quantitive
analysis methods, creation of metabolic images, and
recognition/removal of unwanted artifacts.
|
14:10
|
|
Applications of Whole Organ MRSI - Brain & Body 
Sarah Nelson1
1Radiology and Biomedical Imaging, University
of California, San Francisco, San Francisco, CA, United
States
MR spectroscopic imaging (MRSI) makes it possible to
study changes in metabolism that are associated with
disease progression and response to therapy. Advances in
MR hardware and software have provided new opportunities
for obtaining data in a clinical feasible time and have
therefore opened the door to a much broader range of
applications than was previously considered. These
applications will be demonstrated and future
opportunities described.
|
14:30
|
|
Break & Meet the Teachers |
|
| |
|
|
|
|
| |
|
|
2D MRS |
|
14:50
|
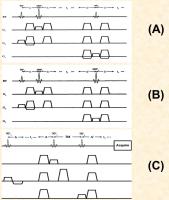 |
Basic Principles & Sequences for 2D MRS 
M. Albert Thomas1, Zohaib Iqbal1,
Manoj Kumar Sarma1, and Rajakumar Nagarajan1
1Radiology, UCLA Geffen School of Medicine,
Los Angeles, CA, United States
In one-dimensional (1D) MR Spectroscopy (MRS), it is
difficult to resolve the multitude of metabolite peaks
that exist over a small spectral range. Spectral-editing
techniques target a particular J-coupled metabolite
selectively, such as lactate, GABA, glutamate, etc. with
a drawback that only one metabolite is selected for each
recording. Due to the added 2nd dimension,
two-dimensional (2D) MRS can unambiguously resolve many
overlapping peaks non-selectively. Instead of a standard
1D spectrum plotting intensity versus a single-axis
(i.e., chemical shift + J-coupling), 2D MRS techniques
produce a 2D spectrum plotting intensity versus two
frequency axes, the dimensions of which depend on the
specific 2D MRS technique. A major goal of this
presentation is to give an overview of the basics of 2D
MRS and describe several localized 2D MRS sequences
which have been implemented on the whole body 1.5T, 3T,
and 7T MRI scanners.
|
15:10
|
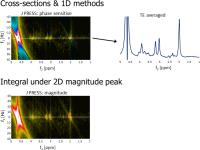 |
Data Analysis for 2D MRS: Spectral Fitting 
Rolf F Schulte1
1GE Global Research, Munich, Germany
Main goal of in vivo Magnetic Resonance Spectroscopy (MRS)
is the determination of individual metabolite
concentrations in organs like the brain. Spectrally
two-dimensional spectroscopy can help to encode more
spectral information during the acquisition, and hence
disentangle the overcrowded proton spectra. In order to
quantify the 2D spectra most accurately, it is necessary
to fit them to 2D metabolite basis spectra, hence
utilising the full amount of available prior
information. Reasons for fitting along with the actual
fitting methods are explained in this educational talk.
|
15:30
|
 |
Applications of 2D MRS: Brain & Body 
Saadallah Ramadan1
1University of Newcastle, Australia
Different types of 2D MRS can offer different types of
information to understand the complexities underlying
pathophysiology of disease. Technical developments
specific to 2D method development and advanced
post-processing methods will allow for the
identification of biomarkers of diseases at an early
stage. Acceleration of signal acquisition, as well as
automated data processing algorithms are essential to
introduce 2D MRS methods into the clinic.
|
15:50
|
|
Break & Meet the Teachers |
|
| |
|
|
|
|
| |
|
|
Spectral Editing |
|
16:10
|
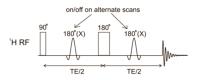 |
Basic Principles and Sequences for Difference and Multiple
Quantum Editing 
Atiyah Yahya1,2
1Department of Oncology, University of
Alberta, Edmonton, AB, Canada, 2Department
of Medical Physics, Cross Cancer Institute, Edmonton,
AB, Canada
In-vivo proton
magnetic resonance spectra exhibit poor spectral
resolution due to the overlap of peaks with similar
chemical shifts. Spectral editing techniques have been
designed and implemented to enable retention of peaks
from metabolites of interest while suppressing
background contaminating peaks. The purpose of the
lecture is to describe two important spectral editing
techniques, namely, difference editing and multiple
quantum filtering. Basic principles and pulse sequences
for each of the methods is presented along with how
spatial localization can be incorporated. In addition,
examples of applications of the sequences are provided.
|
16:30
|
|
Data Analysis for Spectral Editing 
Richard A. E. Edden1
1Russell H Morgan Department of Radiology and
Radiological Science, The Johns Hopkins University
School of Medicine, Baltimore, MD, United States
This presentation will cover the major steps required
for the analysis of edited spectra, which include the
standard steps used for all spectroscopy (Fourier
transformation, windowing/filtering,
integration/fitting) and some steps that are
specifically required by editing (subtraction,
frequency-and-phase correction of time-resolved data).
|
16:50
|
|
Applications of Spectral Editing 
Wolfgang Bogner1
1High-field MR Center, Department of
Biomedical Imaging and Image-guided Therapy, Medical
University Vienna, Vienna, Austria
Performing proton MR spectroscopy (1H-MRS) at
higher static magnetic field strengths (B0)
generally improves spectral resolution and, thereby,
allows detection of a larger number of metabolites.
However, even at very high B0 and
in particular on clinical MR scanners the spectral
resolution is often not sufficient for an unambiguous
quantification of several important J-coupled
metabolites such as GABA, GSH, 2GH, Asc, or Lac. Their
resonances are strongly overlapping with other more
abundant metabolite resonances, which makes their
accurate and reliable quantification via conventional 1H-MRS
difficult. Spectral editing methods can be applied to
selectively quantify these J-coupled metabolites. This
opens the window for numerous clinical and neuro-scientific
applications.
|
17:10
|
|
Roundtable |
17:40
|
|
Adjournment & Meet the
Teachers |
|
| |
|
|
|
|
| |
The International Society for Magnetic Resonance in Medicine is accredited by the Accreditation Council for
Continuing Medical Education to provide continuing medical education for physicians. |




