Oral
Acquisition, Reconstruction & Analysis
Wednesday, 26 April 2017
| Room 311 |
08:15 - 10:15 |
Moderators: Daniel Alexander, Tim Leiner |
Slack Channel: #s_acq_recon_analysis
Session Number: O64
08:15
 |
0685.
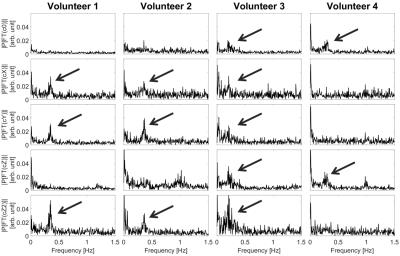 |
Prediction of breathing related B0-field fluctuations via artificial neural networks trained on magnetic field monitoring data 
Niklas Wehkamp, Frederik Testud , Patrick Hucker, Stefan Kroboth, Benjamin Knowles , Jürgen Henning, Maxim Zaitsev
The presented approach utilizes artificial neural networks trained on magnetic field monitoring data in order to predict respiration induced B0-field fluctuations in the brain under the condition of normal breathing. From the predicted B0-field fluctuations it is possible to distinguish the respiration induced resonance offset from the resonance offsets induced by other sources during the course of the experiment. This allows for the quantification of breathing related B0-field fluctuations in the brain of normally breathing healthy volunteers. Furthermore it was observed that the B0-field fluctuations resulting from normal respiration show individual spatial dynamics for every volunteer.
|
08:27
 |
0686.
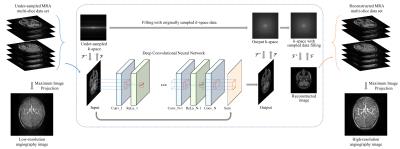 |
Deep Convolutional Neural Network for Acceleration of Magnetic Resonance Angiography (MRA) 
Yohan Jun, Taejoon Eo, Taeseong Kim, Jinseong Jang, Dosik Hwang
In this paper, we propose a deep CNN with skip connection for reconstruction of highly undersampled MRA images. According to the experiments, the proposed method could restore most of fine vessel structures manifested in full-sampled MRA images.
|
08:39
|
0687.
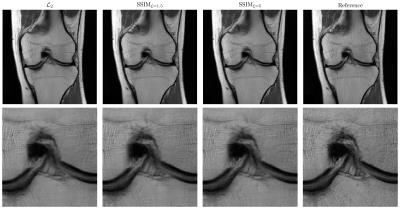 |
L2 or not L2: Impact of Loss Function Design for Deep Learning MRI Reconstruction 
Kerstin Hammernik, Florian Knoll, Daniel Sodickson, Thomas Pock
Human radiologists gain experience from reading numerous MRI images to recognize pathologies and anatomical structures. To integrate this experience into deep learning approaches, two major components are required: We need both a suitable network architecture and a suitable loss function that measures the similarity between the reconstruction and the reference. In this work, we compare pixel-based and patch-based loss functions. We show that it is beneficial to consider other loss functions than the squared L2 norm to get a better representation of the human perceptual system and thus to preserve the texture in the tissue.
|
08:51
|
0688.
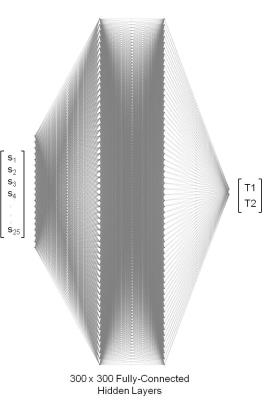 |
Deep learning for fast MR Fingerprinting Reconstruction 
Ouri Cohen, Bo Zhu, Matthew Rosen
The exponential growth in the number of dictionary entries with increasing dictionary dimensions places a practical limit on the number of tissue parameters that may be simultaneously reconstructed. While a sparse sampling of some dimensions can mitigate the problem it also introduces significant errors into the reconstruction. In this work we demonstrate that Deep Learning methods can be used to train a compact neural network with sparse dictionaries without penalty on the reconstruction accuracy.
|
09:03
|
0689.
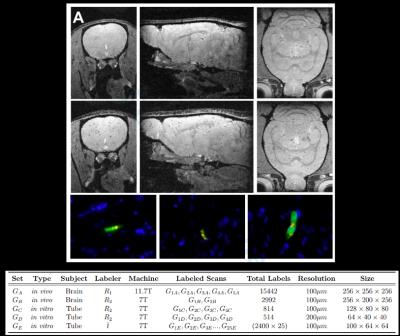 |
Machine Learning for Intelligent Detection and Quantification of Transplanted Cells in MRI 
Muhammad Jamal Afridi, Arun Ross, Erik Shapiro
Cell based therapy (CBT) is promising for treating a number of diseases. The ability to serially and non-invasively measure the number and determine the precise location of cells after delivery would aid both the research and development of CBT and also its clinical implementation. MRI-based cell tracking, employing magnetically labeled cells has been used for the past 20 years to enable detection of transplanted cells, achieving detection limits of individual cells, in vivo.These individual cells can be detected as dark spots in T2* weighted MRI. Manual enumeration of these spots, and hence, counting cells, in an in vivo MRI is a tedious and highly time consuming task that is prone to inconsistency. Therefore, it becomes practically infeasible for an expert to conduct such manual enumeration for a very large scale analysis, consequentially affecting our ability to monitor CBT. To solve this challenge, we have designed a machine learning methodology for automatically quantifying transplanted cells in MRI in an accurate and efficient manner.
|
09:15
|
0690.
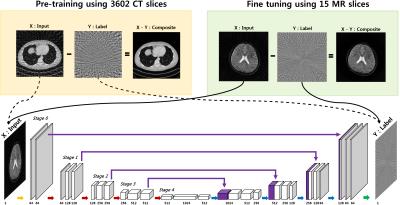 |
Accelerated Projection Reconstruction MR imaging using Deep Residual Learning - permission withheld
Yo Seob Han, Dongwook Lee, JaeJun Yoo, Jong Chul Ye
We propose a novel deep residual learning approach to reconstruct MR images from radial k-space data. We apply a transfer learning scheme that first pre-trains the network using large X-ray CT data set, and then performs a network fine-tuning using only a few MR data set. The proposed network clearly removes the streaking artifact better than other existing compressed sensing algorithm. Moreover, the computational speed is extremely faster than that of compressed sensing MRI.
|
09:27
|
0691.
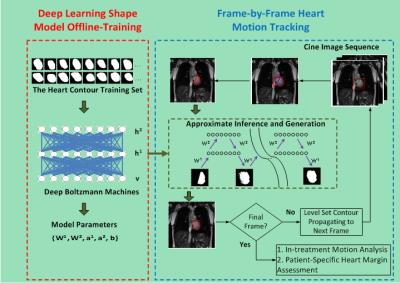 |
Deep Boltzmann Machines-Driven Method for In-treatment Heart Motion Tracking Using Cine MRI 
Jian Wu, Nalini Daniel, Hilary Lashmett, Thomas Mazur, Michael Gach, Laura Ochoa, Imran Zoberi, Su Ruan, Mark Anastasio, Sasa Mutic, Maria Thomas, Hua Li
We developed a hierarchical deep learning shape model-driven method to automatically track the motion of the heart, a complex and highly deformable organ, on two-dimensional cine MRI images. The deep-learning shape model was trained based on a Deep Boltzmann Machine (DBM)1,2 to characterize both global and local shape properties of the heart for accurate heart segmentation on each cine frame. Preliminary experimental results demonstrate the superior shape tracking performance of our proposed method versus two other methods. The tracking method is designed for heart motion pattern analysis during MRI-guided radiotherapy and the subsequent evaluation of potential heart toxicity from radiotherapy.
|
09:39
|
0692.
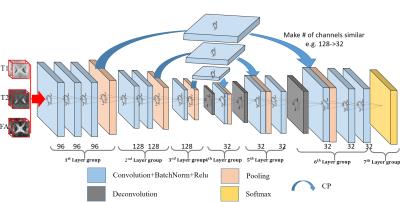 |
Multi-modal Isointense Infant Brain Image Segmentation with Deep Learning based Methods 
Dong Nie, Li Wang, Roger Trullo, Ehsan Adeli, Weili Lin, Dinggang Shen
Accurate segmentation of infant brain images into different regions of interest is one of the most important fundamental steps in studying early brain development. In this paper, we propose a framework based on the recently well-received and prominent deep learning methods. Specifically, we propose a novel 3D multimodal fully convolutional network (FCN) architecture for segmentation of isointense phase brain MR images. Our proposed framework can model the brain tissue structures more accurately compared to the traditional methods. The conducted experiments show that our proposed 3D multimodal FCN model outperforms all previous methods by a large margin, in terms of segmentation accuracy.
|
09:51
|
0693.
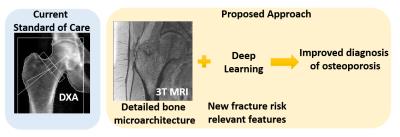 |
Fracture Risk Assessment using Deep Learning and Hip Microarchitecture MRI 
Cem Deniz, Kyunghyun Cho, Stephen Honig, Kenneth Egol, Daniel Sodickson, Gregory Chang
The identification of subjects with high risk of developing osteoporosis-related fracture remains challenging. In this project, we developed supervised convolutional neural networks for hip fracture risk identification using proximal femur MR microarchitecture images and patients’ history of fragility fractures. We found that the proposed fracture risk assessment method provides superior discrimination of fragility fracture patients from controls compared to the current standard of care, DXA.
|
10:03
|
0694.
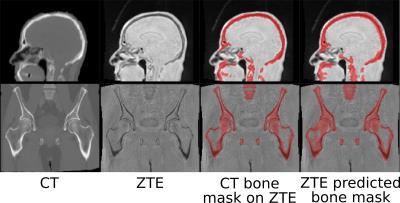 |
Deep Learning based pseudo-CT estimation using ZTE and Dixon MR images for PET attenuation correction 
Sandeep Kaushik, Dattesh Shanbhag, Andrew Leynes, Hariharan Ravishankar, Jaewon Yang, Peder Larson, Thomas Hope, Florian Wiesinger
Simultaneous PET/MR is now being adapted for clinical studies. Earlier methods of PET/MR-AC have considered all bones as a single entity irrespective of their density. In this work, we demonstrate using ZTE and Dixon LAVA-Flex MRI data, and deep learning framework, a continuous density pseudo-CT (pCT) image which combines soft tissue pCT (from Dixon LAVA-Flex) with a continuous density bone pCT from ZTE.
|
|












