Poster: Machine Learning Unleashed
Electronic Power Pitch Poster
Acquisition, Reconstruction & Analysis
14:45 - 15:45
Tuesday, 19 June 2018
Power Pitch Theater A - Exhibition Hall
| |
|
Plasma # |
 |
0428.
 |
1 |
 Deep Learning Method for Non-Cartesian Off-resonance Artifact Correction Deep Learning Method for Non-Cartesian Off-resonance Artifact Correction
David Zeng, Jamil Shaikh, Dwight Nishimura, Shreyas Vasanawala, Joseph Cheng
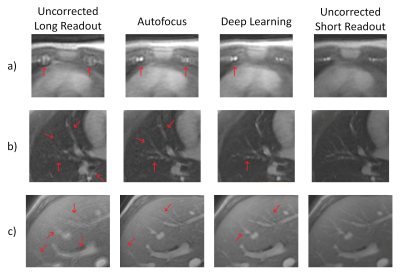
3D cones trajectories have the flexibility to be more scan-time efficient than 3D Cartesian trajectories, especially with long readouts. However, long readouts are subject to blurring from off-resonance, limiting the efficiency. We propose a convolutional residual network to correct for off-resonance artifacts to allow for reduced scan time. Fifteen exams were acquired with both conservative readout durations and readouts 2.4x as long. Long-readout images were corrected with the proposed method. The corrected long-readout images had non-inferior (p<0.01) reader scores in all features examined compared to conservative readout images.
|
|
0429.
 |
2 |
 Gibbs-Ringing Artifact Reduction in MRI via Machine Learning Using Convolutional Neural Network Gibbs-Ringing Artifact Reduction in MRI via Machine Learning Using Convolutional Neural Network
Qianqian Zhang, Guohui Ruan, Wei Yang, Kaixuan Zhao, Ed X. Wu, Yanqiu Feng
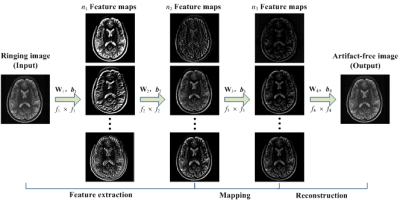
The Gibbs-ringing artifact is caused by the insufficient sampling of the high frequency data. Existing methods generally exploit smooth constraints to reduce intensity oscillations near high-contrast boundaries but at the cost of blurring details. This work presents a convolutional neural network (CNN) method that maps ringing images to their ringing-free counterparts for Gibbs-ringing artifact removal in MRI. The experimental results demonstrate that the proposed method can effectively remove Gibbs-ringing without introducing noticeable blurring.
|
 |
0430.
 |
3 |
 Simultaneous detection and identification of MR artifact types in whole-body imaging Simultaneous detection and identification of MR artifact types in whole-body imaging
Thomas Kuestner, Ke Liu, Annika Liebgott, Lukas Mauch, Petros Martirosian, Fabian Bamberg, Konstantin Nikolaou, Bin Yang, Fritz Schick, Sergios Gatidis
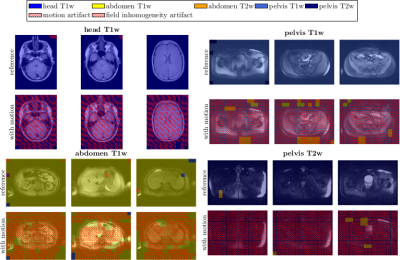
Varying acquisition and reconstruction conditions as well as long examination times make MRI susceptible to various kinds of artifacts. If suitable correction techniques are not available/applicable, if human experts who judge the achieved quality are not present or for epidemiological cohort studies in which a manual quality analysis of the large database is impracticable, an automated detection and identification of these artifacts is of interest. Convolutional neural networks with residual and inception layers localize and identify occurring artifacts. Artifacts (motion and field inhomogeneity) can be precisely identified with an accuracy of 92% in a whole-body setting with varying contrasts.
|
|
0431.
 |
4 |
 Automatic Assessment of MR Image Quality with Deep Learning Automatic Assessment of MR Image Quality with Deep Learning
Jifan Li, Shuo Chen, Qiang Zhang, Huiyu Qiao, Xihai Zhao, Chun Yuan, Rui Li
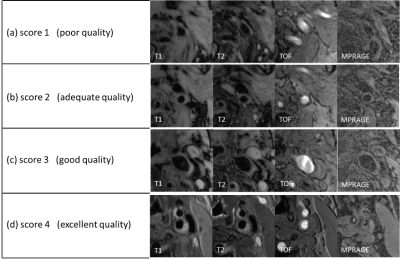
In this study, we aimed to develop a convolutional neural network (CNN) to assess the quality of multi-contrast carotid plaque MR images automatically. The network was trained on large amount of plaque images combined with image quality scores labeled by experienced radiologists. Transfer learning was utilized to take the advantage of state-of-the-art CNN pre-trained on ImageNet dataset. The accuracy of image quality estimation achieved 86.0% with preprocessing and fine-tuning of the network.
|
|
0432.
 |
5 |
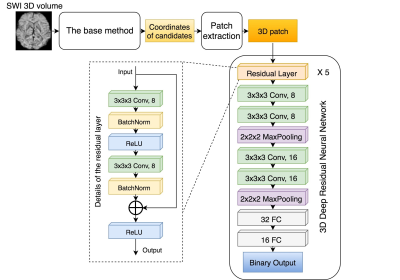
 Automatic detection of cerebral microbleeds using Susceptibility-Weighted Imaging and a 3D deep residual network Automatic detection of cerebral microbleeds using Susceptibility-Weighted Imaging and a 3D deep residual network
Yicheng Chen, Melanie Morrison, Javier Villanueva-Meyer, Janine Lupo
In this abstract, a deep residual neural network based approach that improves the automatic detection and labeling of cerebral microbleeds by significantly reducing the number of false positives compared to previously published algorithms is proposed. This combined method removed 89% of false positives in the test patients with brain tumors who had radiation-induce CMBs while losing only 3% of the true microbleeds and has the potential to fully automate CMB detection.
|
 |
0433.
 |
6 |
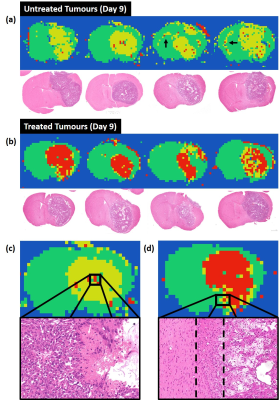
 Deep learning diffusion fingerprinting to detect brain tumour response to chemotherapy Deep learning diffusion fingerprinting to detect brain tumour response to chemotherapy
Thomas Roberts, Ben Hipwell, Giulia Agliardi, Valerie Taylor, Mark Lythgoe, Simon Walker-Samuel
Convolutional neural networks (CNNs) often require very large datasets for robust training and evaluation. As an alternative approach, we introduce deep learning diffusion fingerprinting (DLDF), which treats every voxel as an independent data point, rather than using whole images or patches. We use DLDF to classify diffusion-weighted imaging voxels in a mouse model of glioblastoma, both prior to and in response to Temozolomide chemotherapy. We show that, even with limited training, DLDF can automatically segment brain tumours from normal brain, can distinguish between young and older tumours and that DLDF can detect if a tumour has been treated with chemotherapy.
|
|
0434.
 |
7 |
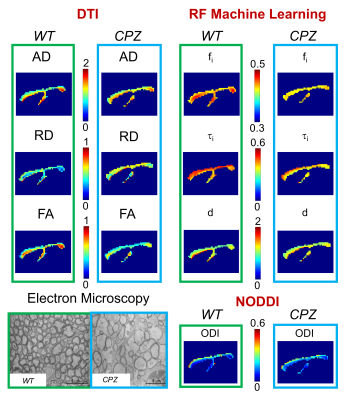
 Machine learning based estimation of axonal permeability: validation on cuprizone treated in-vivo mouse model of axonal demyelination Machine learning based estimation of axonal permeability: validation on cuprizone treated in-vivo mouse model of axonal demyelination
Marco Palombo, Ioana Hill, Mathieu Santin, Francesca Branzoli, Anne-Charlotte Philippe, Demian Wassermann, Marie-Stephane Aigrot, Bruno Stankoff, Hui Zhang, Stephane Lehericy, Alexandra Petiet, Daniel Alexander, Ivana Drobnjak
Estimating axonal permeability reliably is extremely important, however not yet achieved because mathematical models that express its relationship to the MR signal accurately are intractable. Recently introduced machine learning based computational model showed to outperforms previous approximate mathematical models. Here we apply and validate this novel method experimentally on a highly controlled in-vivo mouse model of axonal demyelination, and demonstrate for the first time in practice the power of machine learning as a mechanism to construct complex biophysical models for quantitative MRI.
|
|
0435.
 |
8 |
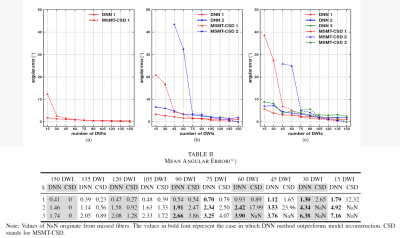
 Direct and Fast Learning of Fiber Orientation Distribution Function for Tractography Direct and Fast Learning of Fiber Orientation Distribution Function for Tractography
Ting Gong, Hongjian He, Zhichao Lin, Zhiwei Li, Qiqi Tong, Yi Sun, Feng Yu, Jianhui Zhong
Multi-shell, multi-tissue, constrained spherical deconvolution is an appealing method for the reconstruction of fiber orientation distribution function (fODF), which is of great importance for solving complex fiber configurations to achieve reliable tractography. However, many diffusion measurements and multiple reconstruction steps are required. In this study, the deep neural network were employed to form a multi-output regression problem for establishing a fast and direct estimation of fODF. The proposed method offers a new streamlined reconstruction procedure which exhibits great potential for accelerating the reconstruction of fODF with whole-brain coverage, with satisfactory accuracy in two minutes.
|
|
0436.
 |
9 |
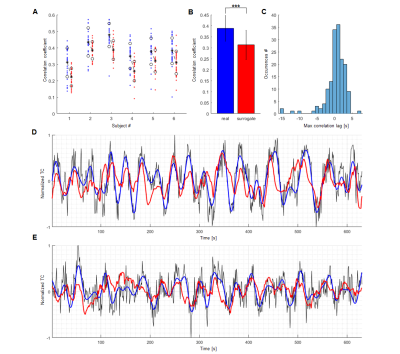
 Predict the slow oscillation of the single-vessel resting-state fMRI signal of rats and humans with echo state networks Predict the slow oscillation of the single-vessel resting-state fMRI signal of rats and humans with echo state networks
Filip Sobczak, Yi He, Xin Yu
Single-vessel fMRI has enabled the detection of slow fluctuations (<0.1Hz) of the hemodynamic fMRI signal from individual vessels in both rat and human brains. The Echo State Network (ESN) has been used to encode the slowly changing temporal dynamics of individual vessels by training the network to predict the oscillatory signals from individual vessels 10 seconds ahead in time. Distinct network reservoirs are optimized for human and animal vascular signals, showing high correlation for the ESN-predictive signal with the original fresh data. This work establishes ESN-based signal prediction for the slow-oscillatory brain fMRI signal in real-time.
|
|
0437.
 |
10 |
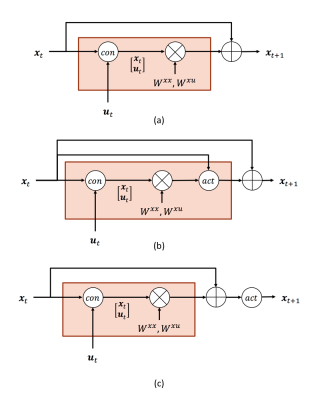
 Dynamic Causal Modelling with neuron firing model in Generalized Recurrent Neural Network framework Dynamic Causal Modelling with neuron firing model in Generalized Recurrent Neural Network framework
Yuan Wang, Yao Wang, Yvonne Lui
DCM-RNN is a Generalized Recurrent Neural Network accommodating the Dynamic Causal Modelling (DCM), which links the biophysical interpretability of DCM and the power of neural networks. It significantly extends the flexibility of DCM, provides unique parameter estimation methods, and offers neural network compatibility. In this abstract, we show how to incorporate neuron firing model into DCM-RNN with ease. An effective connectivity estimation experiment with simulated fMRI data shows that the influence of the firing model is substantial. Ignoring it, as the classical DCM does, can lead to degraded results.
|
 |
0438.
 |
11 |
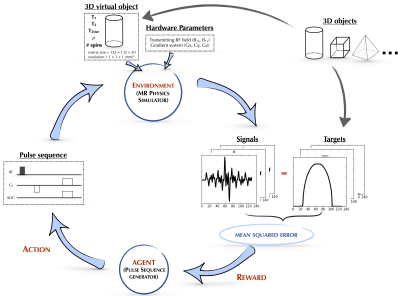
 AUTOmated pulse SEQuence generation (AUTOSEQ) using Bayesian reinforcement learning in an MRI physics simulation environment AUTOmated pulse SEQuence generation (AUTOSEQ) using Bayesian reinforcement learning in an MRI physics simulation environment
Bo Zhu, Jeremiah Liu, Neha Koonjoo, Bruce Rosen, Matthew Rosen
Although the macroscopic equations of motion for nuclear magnetic resonance have been described and modeled for decades by the Bloch equations, limited human intuition of their nonlinear dynamics is an obstacle to fully exploiting the vast parameter space of MR pulse sequences. Here we recast the general problem of pulse sequence development as a game of perfect information, and propose an approach to optimize game play with a Bayesian derivative of reinforcement learning within a MRI physics simulation environment. We demonstrate an AI agent learning a canonical pulse sequence (gradient echo) and generating non-intuitive pulse sequences approximating Fourier spatial encoding.
|
|
0439.
 |
12 |
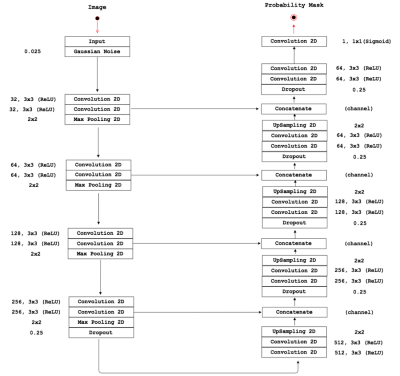
 Towards a fully Automated Time-context Sensitive Convolutional Neural Network for Common Carotid Artery Lumen Segmentation on Dynamic MRI Towards a fully Automated Time-context Sensitive Convolutional Neural Network for Common Carotid Artery Lumen Segmentation on Dynamic MRI
Roberto Souza, Mariana Bento, Lívia Rodrigues, Letícia Rittner, Roberto Lotufo, Richard Frayne
Carotid artery atherosclerosis is one of the main causes of stroke and there is a pressing need for a non-invasive method to quantify, monitor and assess carotid artery stenosis, composition and distensiblity. Here we focus on developing a fully automated convolutional neural network (CNN) with time-context for segmenting the common carotid artery lumen from dynamic magnetic resonance images. The challenge in developing a fully automated carotid segmentation algorithm is that there are other vessels with size and spatial location comparable to the carotid artery. Our preliminary results indicate that a CNN with time-context is capable of distinguishing and segmenting the carotid artery from other vessels.
|
|
0440.
 |
13 |
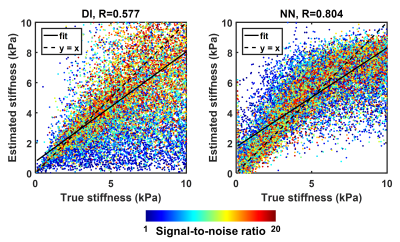
 Artificial neural networks for stiffness estimation in magnetic resonance elastography Artificial neural networks for stiffness estimation in magnetic resonance elastography
Matthew Murphy, Armando Manduca, Joshua Trzasko, Kevin Glaser, John Huston, Richard Ehman
Artificial neural networks (ANNs) were trained using simulated displacement fields to perform stiffness estimation from MRE data. These neural network-based inversions (NNIs) are evaluated in simulation and in vivo. In a test set of simulated data, NNI is shown to provide a more accurate estimate of stiffness compared to a standard direct inversion (DI) approach. In vivo, NNI-based stiffness strongly correlated with DI-based stiffness across a range of fibrosis stages in the liver and ages in the brain, indicating that NNI can detect relevant biology. Finally, test-retest error in the brain is reduced using NNI compared to DI.
|
|
0441.
 |
14 |
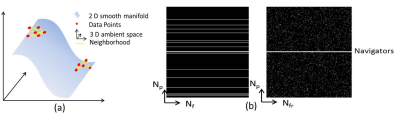
 MLS: Self-learned joint manifold geometry and sparsity aware framework for highly accelerated cardiac cine imaging MLS: Self-learned joint manifold geometry and sparsity aware framework for highly accelerated cardiac cine imaging
Ukash Nakarmi, Konstantinos Slavakis, Hongyu Li, Chaoyi Zhang, Peizhou Huang, Sunil Gaire, Leslie Ying
In this work, we propose a novel joint manifold learning and sparsity aware framework for highly accelerated cardiac cine imaging. The proposed method efficiently captures the intrinsic low dimensional nonlinear manifold geometry and inherent periodicity of cardiac data, and outperforms the current state-of-the-art accelerated MRI methods.
|
|
0442.
 |
15 |
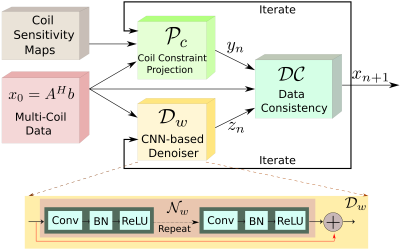
 MoDL: Model Based Deep Learning Architecture for Image Recovery with Prior Information. MoDL: Model Based Deep Learning Architecture for Image Recovery with Prior Information.
Hemant Kumar Aggarwal, Merry Mani, Mathews Jacob
The primary focus of this work is to introduce a novel deep learning framework, which synergistically combines the benefits of model-based image recovery with the power of deep learning. This work enables the easy exploitation of prior information available from calibration scans, in addition to significantly reducing the number of network parameters, amount of training data required, and computational complexity. More importantly, the insensitivity of the learned model to the acquisition parameters also facilitates its easy reuse with a range of acquisition settings.
|
|

 Watch the full Pitch Session Here
Watch the full Pitch Session Here














