Single-Voxel MRS Techniques
Single-Voxel MRS Techniques
Oral
Oral
Spectroscopy & Non-Proton MR
Thursday, 16 May 2019
| Room 513D-F | 08:15 - 10:15 | Moderators: Ariane Fillmer, Arend Heerschap |
08:15 |
1060. 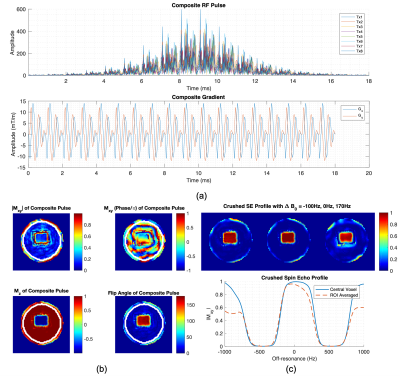 |
Parallel Transmit Optimized Spiral Composite Adiabatic Pulses for 3D Spatial-Spectral Selectivity in Spectroscopy
Xiaoxuan He, Edward Auerbach, Michael Garwood, Xiaoping Wu, Gregory Metzger
In this abstract we demonstrated by simulation and phantom experiments the feasibility of designing composite refocusing pulse for spectroscopy using parallel transmission, featuring inherent 2D spatial and spectral selectivity along with improved immunity to B0 non-uniformity and B1+ heterogeneity while simultaneously providing lipid and water suppression and short echo times of ~18ms.
|
08:27 |
1061. 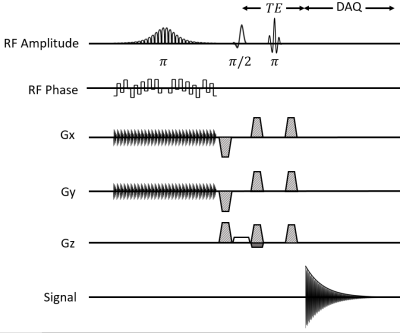 |
Double-Shot Semi-Adiabatic Localization for Ultra Short Echo Time Magnetic Resonance Spectroscopy
Karl Landheer, Michael Garwood, Ralph Noeske, Christoph Juchem
Modern magnetic resonance spectroscopic (MRS) pulse sequences which employ localization through adiabatic pulses suffer from long minimum echo times ( 19
ms). Here we developed a novel MRS pulse sequence which allows for an echo time of 8 ms with adiabatic localization in two of the three spatial dimensions, at the cost of two-shot volume localization. The sequence is highly robust to B1 inhomogeneity and does not have any anomalous J-modulation since no slice-selective refocusing pulses are used. High spectral quality was obtained in phantom experiments, although strong lipid contamination was observed, consistent with previously reported two-shot localization methods.
|
08:39 |
1062. 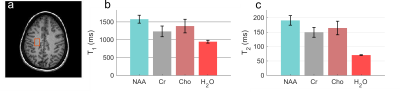 |
Optimization of 1H Single Voxel Multi-Parametric Spectroscopic Fingerprinting (MRSF)
Alexey Onikul Kulpanovich , Assaf Tal
Brain metabolite T1 and T2 have diagnostic value and per-subject knowledge of both metabolite and water reference T1, T2 is necessary for accurate metabolite absolute quantification. Here we optimize the excitation schedules of our recently-proposed magnetic resonance spectroscopic fingerprinting (MRSF) sequence to accurately estimate metabolite T1 and T2. We computationally evaluate the optimized schedules using Monte-Carlo simulations, as well as phantom and in-vivo experiments.
|
| 08:51 |
1063. 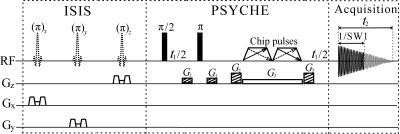 |
Localized pure shift proton MRS for measurements on biological samples
Yuqing Huang, Jin Yan, Chunhua Tan, Zhong Chen
Localized proton magnetic resonance spectroscopy (MRS) provides a non-invasive tool for metabolic studies on biological systems by revealing valuable biochemical information, complementary to anatomical information delivered by MRI. However, due to limited proton chemical shift range and extensive J coupling splittings, spectral congestion is generally encountered in the resulting 1D spectra acquired by conventional proton MRS techniques, such as PRESS (Point resolved spectroscopy) and STEAM (STimulated Echo Acquisition Mode). In this abstract, we present a previously-unreported proton MAS approach to obtain localized 1D pure shift spectra with spectral simplification, potentially useful for studies on biological samples.
|
09:03 |
1064 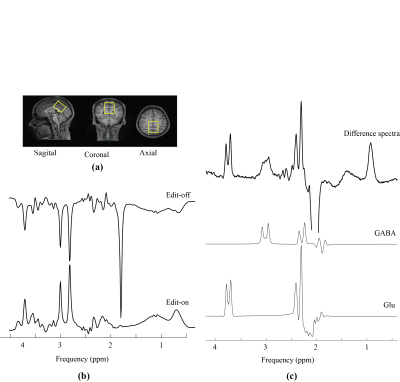 |
In vivo Detection of Metabolic Turnover of GABA and Glutamate in Human Brain using Dynamically Acquired MEGA-PRESS MRS During 13C-Labeled Glucose Infusion Video Permission Withheld
Masoumeh Dehghani, Pierre Etienne, Jamie Near
Glucose, the main substrate for cerebral energy metabolism, serves as a metabolic precursor for both glutamate and GABA synthesis. In the current study, we employ a novel approach to investigate 13C-labeling of both glutamate and GABA in the human brain. Specifically, localized homonuclear (1H) J-difference edited (MEGA-PRESS) MRS spectra were acquired dynamically (without heteronuclear decoupling or editing pulses) to detect glutamate and GABA labelling following an infusion of 13C labeled glucose. Despite excellent spectral quality and temporal stability, little or no GABA labeling was observed, raising some questions as to the functional status of the GABA pools detected by MEGA-PRESS.
|
09:15 |
1065. 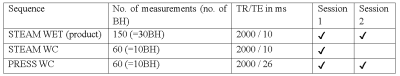 |
Cardiac creatine quantified by cycled water-suppressed 1H-MRS at 3T
Belinda Ding, Mark Peterzan, Ferenc Mózes, Hannah Lake, Craig Lygate, Rana Sayeed, Mario Petrou, George Krasopoulos, Oliver Rider, Ladislav Valkovic, Christopher Rodgers
This work aimed to use cardiac 3T 1H-MRS to quantify creatine concentration (“[Cr]”) in the human heart, because creatine plays a key role in cardiac energy metabolism. We implemented a cycled water-suppression scheme (WC) into the PRESS and STEAM pulse sequences on a Siemens 3T Prisma MRI scanner. Compared to the vendor’s product PRESS sequence, we found that PRESS-WC was able to measure [Cr] more than 2 times more reproducibly and in a shorter scan. Using PRESS WC in a 10-breathhold protocol, we were able to detect a decrease in creatine concentration in 5 patients suffering from aortic stenosis.
|
09:27 |
1066. 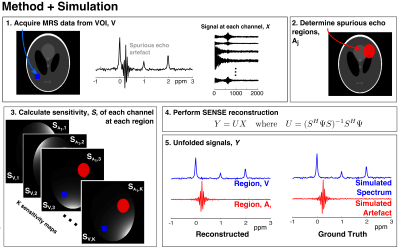 |
SENSE-based reconstruction for removal of spurious echo artifacts in MRS
Adam Berrington, Michal Povazan, Peter Barker
Spurious echo artifacts appear in MRS spectra and originate from regions of unsuppressed and insufficiently crushed signal. We propose a reconstruction method to remove such artifacts using the sensitivity weighting of the receive channels at each spatial location. Data from phantom and simulations show that spurious echoes can be separated from underlying signal. The spatial distribution of unwanted signal was estimated in vivo using B0 maps. Artifactual components of the signal were identified in sinus regions, however not entirely removed from the spectrum. Further work will aim to improve the conditioning of the SENSE reconstruction to remove spurious echo artifacts from MRS data.
|
09:39 |
1067. 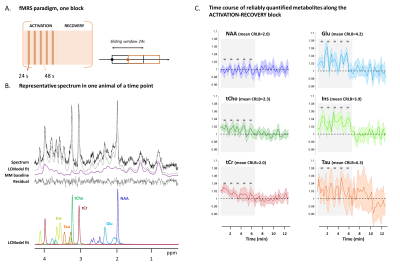 |
Functional Magnetic Resonance Spectroscopy in the mouse brain
Clémence Ligneul, Julia Huntenburg, Francisca Fernandes, Noam Shemesh
Functional magnetic resonance spectroscopy (fMRS) can provide insights on brain metabolism under activation, as has already been shown in humans. Assessing fMRS in mice would open interesting perspectives for understanding neural activity given mouse transgenics. We report preliminary encouraging results of fMRS in the mouse superior colliculus, during visual stimulation at 9.4 T (using a cryocoil for reception). Time courses of different metabolites concentrations notably reveal metabolite signal modulations during activation.
|
09:51 |
1068. 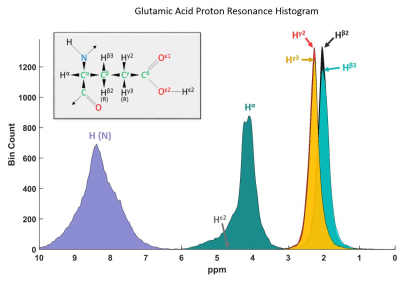 |
Towards a Fitting Model of Macromolecular Spectra: Amino Acids
Tamás Borbáth, Saipavitra Murali Manohar, Anke Henning
Broad signals underlying in vivo 1H MRS spectra are referred to in
|
10:03 |
1069. 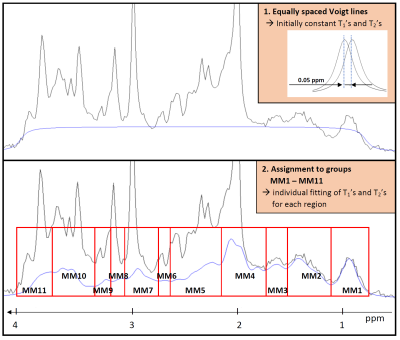 |
About the need for a comprehensive description of the macromolecular baseline signal for MR fingerprinting and multidimensional fitting of MR spectra
Maike Hoefemann, Christine Bolliger, Jan Willem van der Veen, Roland Kreis
The purpose of this study was to evaluate the need for a comprehensive description of the macromolecular baseline signal (MMBL), e.g. for MR fingerprinting and multidimensional fitting of MR spectra. Inherent J-evolution for certain signals as well as multi-component T2-decays can cause variance of the signal shape as function of TE that is not described in a mono-exponential decay model. Two methods are described to accommodate this signal behavior when fitting a 2DJ-Inversion-Recovery dataset. In addition, the MMBLs for different TEs, as well as resulting metabolite contents, T1s and T2s with the conventional and alternative methods are reported.
|
 Back to Program-at-a-Glance |
Back to Program-at-a-Glance |  Back to Top
Back to Top